 With the help of WGCNA, a set of four long non-coding RNAs (lncRNAs) were found to be significantly correlated with the carcinogenesis and progression of colon adenocarcinoma (COAD) ( Jiang et al., 2019). et al., 2020), while FBXW4 was reported to be associated with chemotherapy resistance and prognosis of CRC ( Zhang Y. 
In terms of CRC, PIGU was identified as a key modulator that is closely related to KRAS mutant CRC patients ( Zhang M. In recent years, WGCNA has been widely used to uncover the potential biomarkers associated with clinical parameters in various cancer types ( Liang et al., 2020 Wang et al., 2020). Compared with the other methodologies applied to analyze the high-throughput sequencing data, WGCNA implements methods for both weighted and unweighted correlation networks and provides a more effective mean to explore the potential association between modules and sample traits. By utilizing the WGCNA algorithm, genes with similar co-expression patterns are classified into a set of modules, in which the most central genes could be further identified as hub genes. Among them, the weighted gene co-expression network analysis (WGCNA) approach provides a systematic biology strategy to identify modules of highly correlated genes and construct a co-expression gene network ( Langfelder and Horvath, 2008). With the recent developments in bioinformatics, a number of well-designed and effective methods are now available for the comprehensive identification of biomarkers in cancer and the prediction of cancer-related signaling pathways ( Carter et al., 2004 Carey et al., 2005 Liu et al., 2007). Therefore, identifying the key regulators involved in CRC recurrence and deciphering their underlying mechanisms are critical for CRC prognosis and the development of novel therapeutic drugs. Despite the massive progress in understanding its genetic mechanism and improvement of surgical techniques, the overall survival time of CRC patients is still not remarkably improved, which could be ascribed primarily to the high recurrence rates ( Yu et al., 2017 Ha et al., 2019 Mirgayazova et al., 2019). In conclusion, the identification of these hub genes and the prediction of the Regulatory relationship of TFs-miRNAs-hub genes may provide a novel insight for understanding the underlying mechanism for CRC recurrence.Ĭolorectal cancer (CRC) ranks as the third most common type of tumor around the world and accounts for more than 7% of overall cancer-related death in China ( Liu et al., 2015 Siegel et al., 2020). 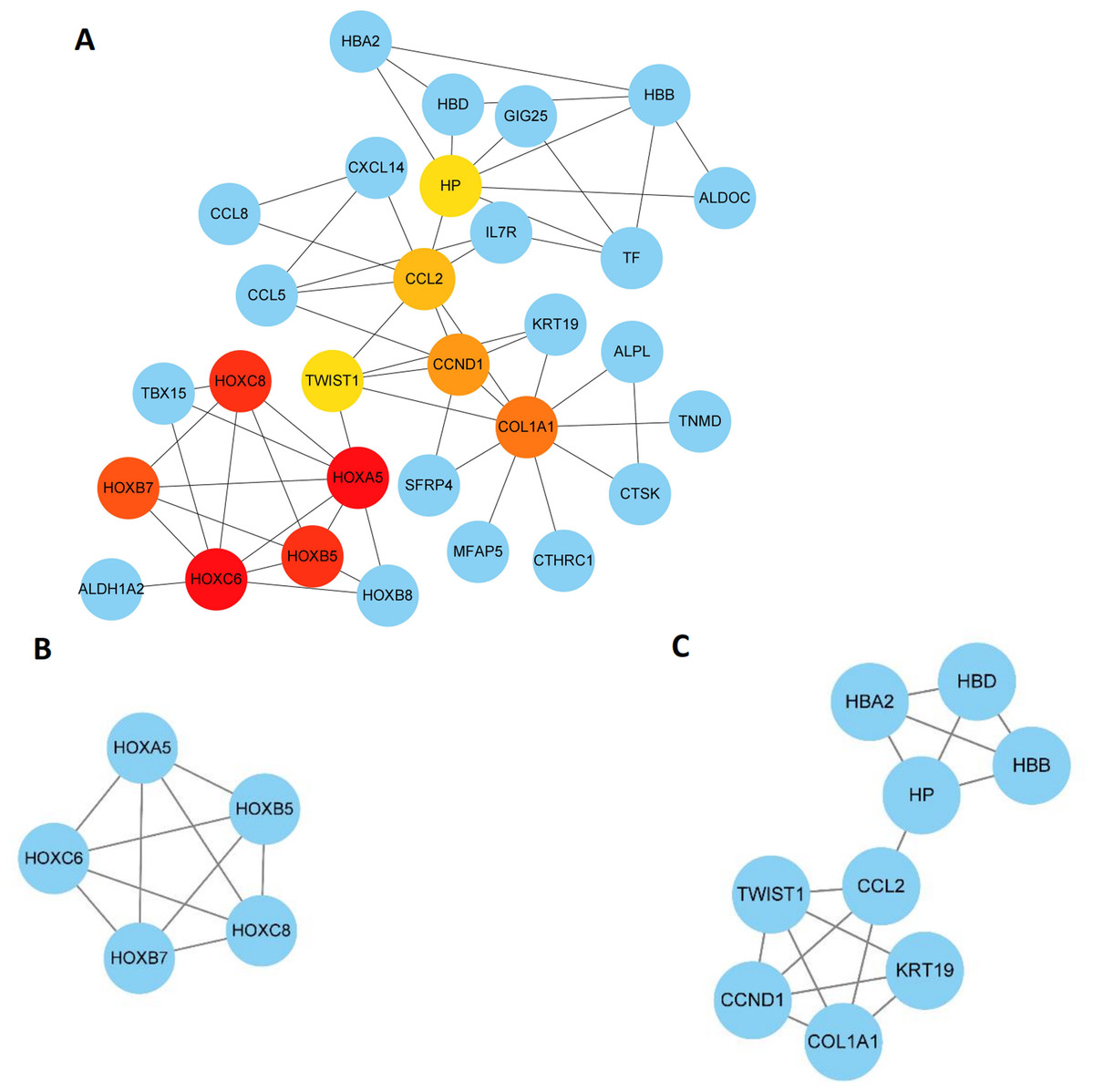
A regulatory network of TFs-miRNAs-targets with 29 TFs, 58 miRNAs, and five hub genes was instituted, including model GATA6-MIR106A-CNN1, SP4-MIR424-TPM2, SP4-MIR326-MYL9, ETS1-MIR22-TIMP1, and ETS1-MIR22-SPARCL1. After identification and validation, a total of five genes (TIMP1, SPARCL1, MYL9, TPM2, and CNN1) were selected as hub genes. Construction of a TFs-miRNAs–hub genes network was also conducted using StarBase and Cytoscape approaches. Samples from the Cancer Genome Atlas (TCGA) were further used to validate the hub genes. The protein and protein interaction network was then built to screen hub genes. 
Functional annotation of the key module genes was assessed through Gene Ontology and Kyoto Encyclopedia of Genes and Genomes analyses. A total of 177 cases from the GSE17536 dataset were analyzed via weighted gene co-expression network analysis to explore the modules related to CRC recurrence. This study aimed to construct a gene co-expression network to predict the hub genes affecting CRC recurrence and to inspect the regulatory network of hub genes and transcription factors (TFs). However, the key regulators dictating this process and their exact mechanisms are understudied. Tumor recurrence is one of the most important risk factors that can negatively affect the survival rate of colorectal cancer (CRC) patients. 2Chongqing Key Laboratory of Biochemistry and Molecular Pharmacology, School of Pharmacy, Chongqing Medical University, Chongqing, China.1Department of Pharmacy, Yongchuan Hospital of Chongqing Medical University, Chongqing, China.Shengwei Liu 1,2, Fanping Zeng 1, Guangwen Fan 1 and Qiyong Dong 1* 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
